Flexible Purification Solutions from Process Development to Commercialization
Our industry-leading bioseparation technologies empower our customers to deliver life-improving and life-saving advanced therapies to patients worldwide. With a legacy of over 30 years, our team takes pride in supplying world-class products that adhere to FDA and EMA standards, ensuring excellence in manufacturing processes.
At the heart of our commitment is a dedicated R&D team relentlessly driving innovation, bringing state-of-the-art technologies to market. In an ever-evolving landscape, we strive to meet the dynamic needs for speed and efficiency. Explore the pinnacle of bioseparation excellence with Astrea Bioseparations, a Biotage company, where every breakthrough propels us towards a future of enhanced patient care.
Customer Learning & Support
Explore the depth of our expertise and enhance your understanding of our cutting-edge technologies by delving into our resources section.
Uncover a wealth of knowledge through our meticulously crafted Brochures, User Guides, Application Notes, Posters, and more. Whether you're a seasoned researcher or a newcomer to the field, our comprehensive collection is designed to empower you with insights that push boundaries.
Explore Resources
What do our customers have to say?
Featured
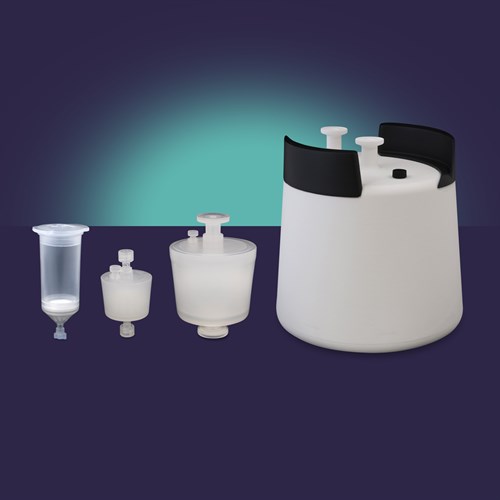
Lentiviral vector purification tools, for high recovery of LVV.
Select product for size specific features, benefits, and technical data.
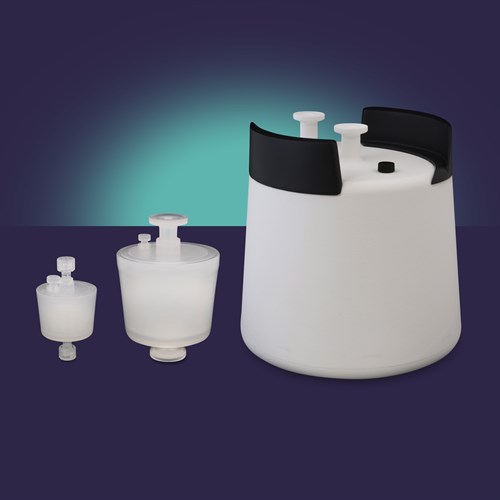
Extracellular vesicle isolation capsule for high recovery of EVs, including exosomes, 1 mL and 10 mL bed volume.
Select product for size specific features, benefits, and technical data.
Streamline pDNA processing with high-speed performance, low operational pressure, and seamless scalability to GMP manufacturing. Ideal for primary capture or polishing steps, including the purification of linearized pDNA.
Select product for size specific features, benefits, and technical data.
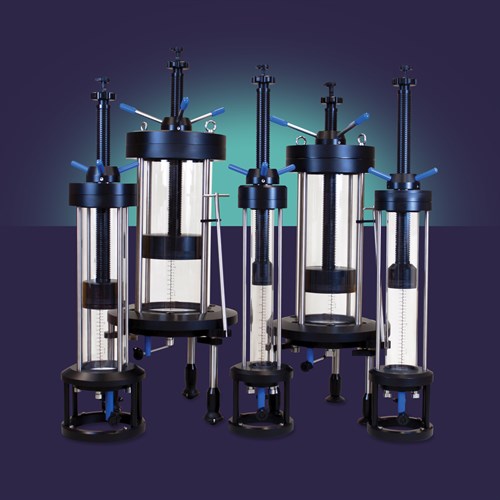
Evolve® Process Columns have been designed specifically for use in biopharmaceutical applications. Available in eight diameters, Evolve® Process Columns provide cost effective, scalable, high performance technology suitable for use in both pilot and production scale chromatographic processes.
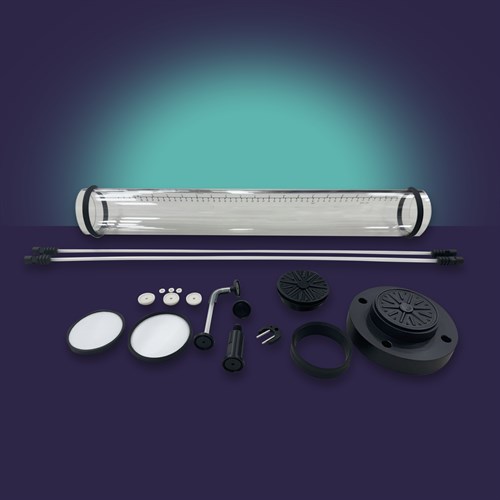
Optimize your chromatographic processes with our Evolve® Refresh Kits, designed for quick, single-use applications and to prevent cross-contamination in all Evolve® Process Columns.
HCPure™ is a mixed mode resin designed with the removal of Host Cell Proteins (HCP) in mind. With a unique operational space compared to other commercially available products, HCPure™ can provide alternative strategies for the removal of problematic HCPs and other process impurities.
An advanced endotoxin removal adsorbent with a high affinity for endotoxins, enabling the efficient removal of up to 99.9% while preserving protein yield. Designed with a high-capacity synthetic ligand, it ensures optimal endotoxin purification and reliable performance in downstream processing across a range of target proteins.